CCA1 mRNA accumulates around dawn and thus represses the expression of its evening-phased targets ( 13, 18). Primarily, CCA1 functions as a transcriptional repressor that binds to the evening element (EE), a motif enriched in promoters of genes with peak expression in the evening ( 3, 13, 20– 22). CCA1, along with LHY and 11 other members of REVEILLE (RVE) proteins, belongs to a subfamily of MYB-domain–containing transcription factors. Together these three components contribute to the rhythmic expression of several other circadian regulators and output genes ( 17– 19). The circadian clock in plants involves a posttranscriptional component and a well-studied transcriptional–translation feedback regulation between the Myb-like transcription factors CIRCADIAN CLOCK ASSOCIATED 1 ( CCA1) and LATE ELONGATED HYPOCOTYL ( LHY) and a member of the PSEUDO RESPONSE REGULATOR ( PRR) family, TIMING OF CAB2 EXPRESSION 1 ( TOC1) ( 10– 16). Together, our results emphasize an expanded role for the clock in regulating a diverse category of genes and key pathways in Arabidopsis and provide a comprehensive resource for future functional studies. Furthermore, this work revealed several CCA1 targets that do not cycle in either LL or LD conditions. Although many of these target genes are evening expressed and contain the EE motif, a significant subset is morning phased and enriched for previously unrecognized motifs associated with CCA1 function. CCA1 targets are enriched for a myriad of biological processes and stress responses, providing direct links to clock-controlled pathways and suggesting that CCA1 plays an important role in regulating a large subset of the rhythmic transcriptome. In this study, using ChIP followed by deep sequencing of CCA1 in constant light (LL) and diel (LD) conditions, more than 1,000 genomic regions occupied by CCA1 were identified. However, the precise molecular mechanisms underlying clock regulation of the rhythmic transcriptome, specifically how clock components connect to clock output pathways, is poorly understood. As a key component of the clock, the Myb-like transcription factor CIRCADIAN CLOCK ASSOCIATED1 ( CCA1) is able to initiate and set the phase of clock-controlled rhythms and has been shown to regulate gene expression by binding directly to the evening element (EE) motif found in target gene promoters. + CpG Islands, repeats, etc.The circadian clock in Arabidopsis exerts a critical role in timing multiple biological processes and stress responses through the regulation of up to 80% of the transcriptome. 
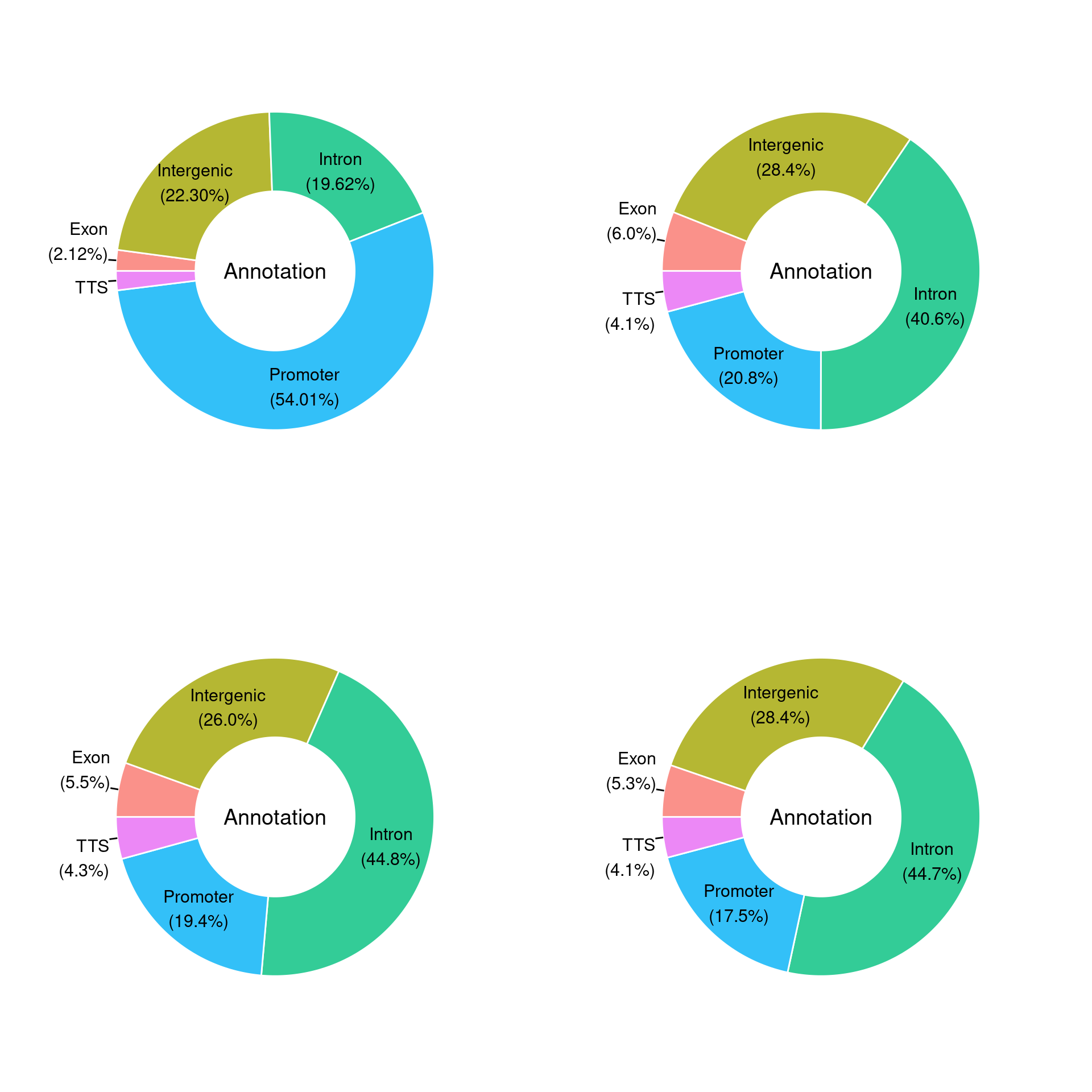
The ChIPSeq or ChIPExo peaks were annotated to regions in the genome () using Homer. All peaks are observed at L4, identifies whether that specific peak is also observed at L8, L10, L15 or L20. By studying the data with respect to a multiple spectrum of peak selection criteria, we adjust for both the risk of excess background noise and the risk of filtering out any low-amplitude information. This performs a low-to-high stringency analysis of the data. Multiple peak selection criteria were used (L4, L8, L10, L15, L20), where Lx represents an x-fold greater tag density at peaks than in the surrounding 10-kb region. We overlapped the location coordinates of the 81,922 0-nt to 5-nt variant 13-nt ERE or HRE DNA elements in the genome and the location coordinates of the ChIPSeq or ChIPExo peaks in an experiment (157 ER experiments at 0-nt to 5-nt variant ERE DNA elements) (194 KR experiments at 0-nt to 5-nt variant HRE DNA elements) to determine the absolute number of times each 0-nt to 5-nt variant 13-nt ERE or HRE DNA element occurred within an experiment (the entire 13-nt DNA element was required to be within the peak boundaries).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
